by Julianne Audette and Coralie Reymond
Introduction
May 19, 2011, the Robert Koch Institute (RKI) in Germany was notified of a number of cases of bloody diarrhea in northern Germany. The RKI is Germany’s public health institute that investigates all types of outbreaks. These cases were soon identified to be caused by the O104:H4 strain of Shiga toxin-producing Escherichia coli (STEC) bacteria and they rapidly evolved into a full-blown outbreak. Ultimately, almost 4,000 people in 16 countries were affected, but the Northern part of Germany was the most affected, as you can see from Figure 1. The outbreak lasted for approximately two months, from the beginning of May to the beginning of July, with a peak during the month of May.
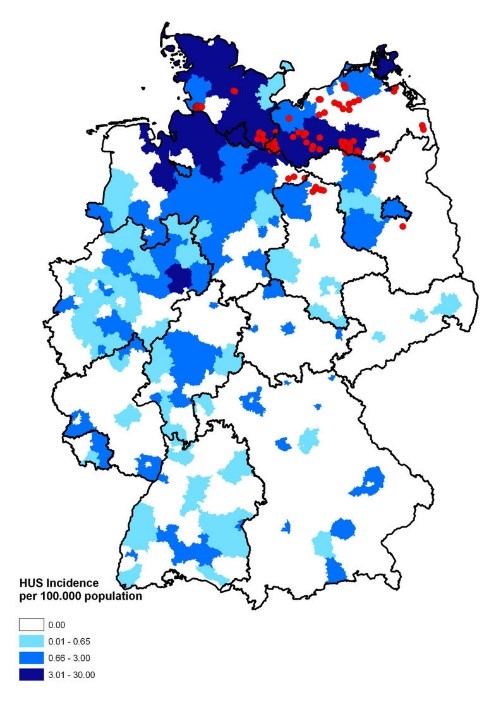 Figure 1: Map of Germany displaying the incidence of HUS [hemolytic uremic syndrome] cases. Source: Wieler LH, Torsten S, Eichhorn I, Antao EM, Kinneman B, Geue L, Karch H, Guenther S, Bethe A. 2011. No evidence of the Shiga toxin-producing E. coli O104:H4 outbreak strain or enteroaggregative E. coli (EAEC) found in cattle faeces in northern Germany, the hotspot of the 2011 HUS outbreak area. Gut Pathogens. [updated 3 November 2011; accessed 17 November 2017]; https://gutpathogens.biomedcentral.com/articles/10.1186/1757-4749-3-17. doi: 10.1186/1757-4749-3-17
Figure 1: Map of Germany displaying the incidence of HUS [hemolytic uremic syndrome] cases. Source: Wieler LH, Torsten S, Eichhorn I, Antao EM, Kinneman B, Geue L, Karch H, Guenther S, Bethe A. 2011. No evidence of the Shiga toxin-producing E. coli O104:H4 outbreak strain or enteroaggregative E. coli (EAEC) found in cattle faeces in northern Germany, the hotspot of the 2011 HUS outbreak area. Gut Pathogens. [updated 3 November 2011; accessed 17 November 2017]; https://gutpathogens.biomedcentral.com/articles/10.1186/1757-4749-3-17. doi: 10.1186/1757-4749-3-17
Disease description
Escherichia coli is a gram negative, rod-shaped bacteria, see Figure 2, which is also a facultative anaerobe. Most of this bacteria’s strains are commensals and reside in gastrointestinal tract as part of the normal microbiota without causing harm to the host. However, different strains of Escherichia coli have acquired virulence factors that render them pathogenic, capable of causing disease. They acquired virulence factors through mutation or horizontal gene transfer, genes from other bacteria. Escherichia coli is one of the most important food-borne pathogens. Enterohemorrhagic Escherichia coli (EHEC) produce the Shiga toxin which causes hemorrhagic colitis (HC) and hemolytic uremic syndrome (HUS). HC and HUS can cause failure of essential organs, such as kidney and heart, leading to death if left untreated. Those bacteria are highly infectious because they survive for a long period of time outside of the host which increases the time products stay contaminated and increases chances of contracting the infection. Also, they can survive extreme conditions, such as the acidic environment of the stomach. Cases related to Escherichia coli are lower than other bacteria types (Salmonella or Campylobacter spp.), however much higher mortality and hospitalization rates are related to them. Escherichia coli’s natural reservoir is in cattle and the farm environment. Cattle are asymptomatic carriers and shed this bacteria in their feces.
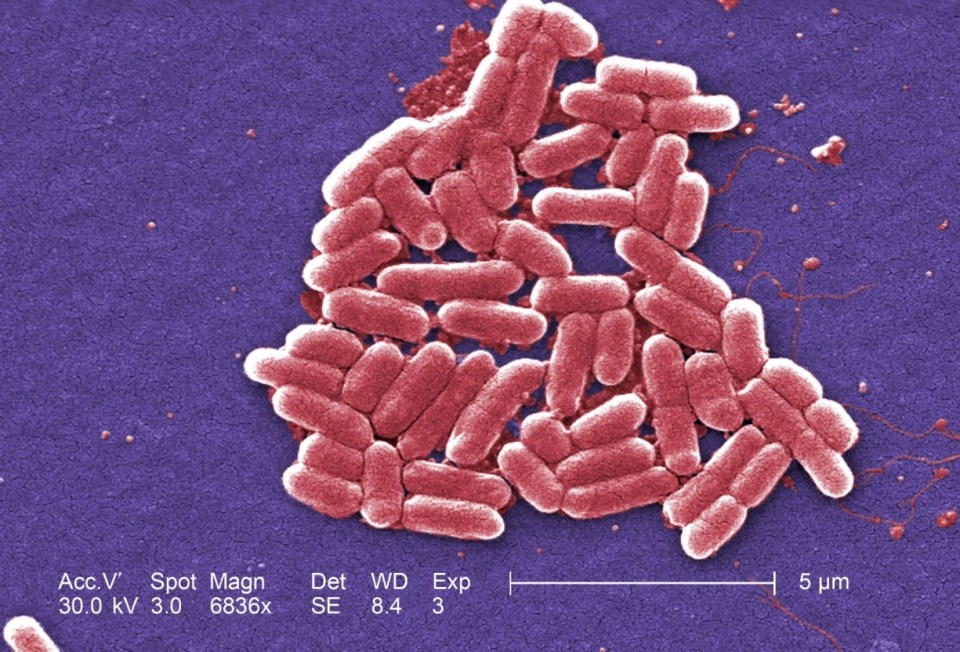 Figure 2: Under a high magnification of 6836X, this digitally-colorized, scanning electron microscopic (SEM) image depicted a growing cluster of Gram-negative, rod-shaped, Escherichia coli bacteria of the strain O157:H7, which is a pathogenic strain of E. coli. Source: Public Health Image Library, Center for Disease Control, Dr. Janice Carr (2006).
Figure 2: Under a high magnification of 6836X, this digitally-colorized, scanning electron microscopic (SEM) image depicted a growing cluster of Gram-negative, rod-shaped, Escherichia coli bacteria of the strain O157:H7, which is a pathogenic strain of E. coli. Source: Public Health Image Library, Center for Disease Control, Dr. Janice Carr (2006).
Discussion of the source of the outbreak
The outbreak was mainly centered in northern Germany and spread throughout Europe, but very little in the United States, as only 6 cases of people having traveled to this area in Germany were recorded. The common factor of all the people infected was that they ate raw vegetables grown locally in northern Germany. The Robert Koch Institute was able to narrow down the source of the outbreak to raw sprouts. Other cases showed up near Bordeaux, France where the infected people had eaten raw sprouts grown locally. This information led the European Food Safety Authority to discover the real source was from fenugreek seeds imported from Egypt into France and Germany. The origin of the outbreak strain and how the seeds were contaminated remains unclear to this day.
Cause of the outbreak
This outbreak was caused by the O104:H4 strain of Escherichia coli rather than the most common strain O157:H7. The high number of HUS cases and deaths indicated that the O104:H4 strain had an augmented virulence compared to the common strain. The exact reason remains unknown. However, scientists have offered various hypotheses. The outbreak strain combines virulence potentials of STEC and enteroaggregative E. coli (EAEC). The distinguishing feature of EAEC is their ability to attach to tissue cells. It has been hypothesized that O104:H4 is a typical EAEC strain that acquired the stx2a gene that encodes for the toxin (mutant). This toxin enters the epithelium cells that line the gastrointestinal tract and lyses the cells which causes bloody diarrhea and HUS. This outbreak pathogen had a longer incubation period (8 days) compared to the O157:H7 strain (3-4 days) which means that there was a longer period between the exposition to the pathogen and the onset of the symptoms, making it more difficult to find the source. The O104:H4 strain also had an increased adherence to intestinal epithelial cells which might facilitate systemic absorption of the toxin. This could explain the high number of patients developing HUS. Antibiotic resistance could also have played a role in this outbreak. This strain is resistant to β-lactam drugs and if this type of antibiotic was used to treat the infection, it would only suppress the competing microbiota making it easier for the O104:H4 strain of Escherichia coli to proliferate.
Measure taken to end the outbreak
To counteract the disease, the infected patients required hospitalization to maintain fluid and electrolyte levels, to monitor and support kidney function and dialysis in case of renal failure. During the outbreak, public health reviewed the potential therapies going above and beyond symptomatic treatment. Two avenues were explored: antibiotic treatment and passive immunity (toxin neutralizing antibodies). Antibiotic therapy was considered only in very severe cases due to increased release of toxin by bacteria lysis. The antibody treatment was not feasible as they were not yet approved meaning that they were still in the clinical research phase.
Authorities were able to identify the origin at the same time as the epidemic curve was starting to decrease. This decline is due to self-limitation, meaning that there was limited exposure to contaminated food in circulation (Figure 3). Raw sprouts were identified as the outbreak source in less than 3 weeks which is relatively rapid and comparable to other outbreak investigations in other countries (incl. US). On July 5th, the seeds from Egypt were banned which definitively prevented further cases.
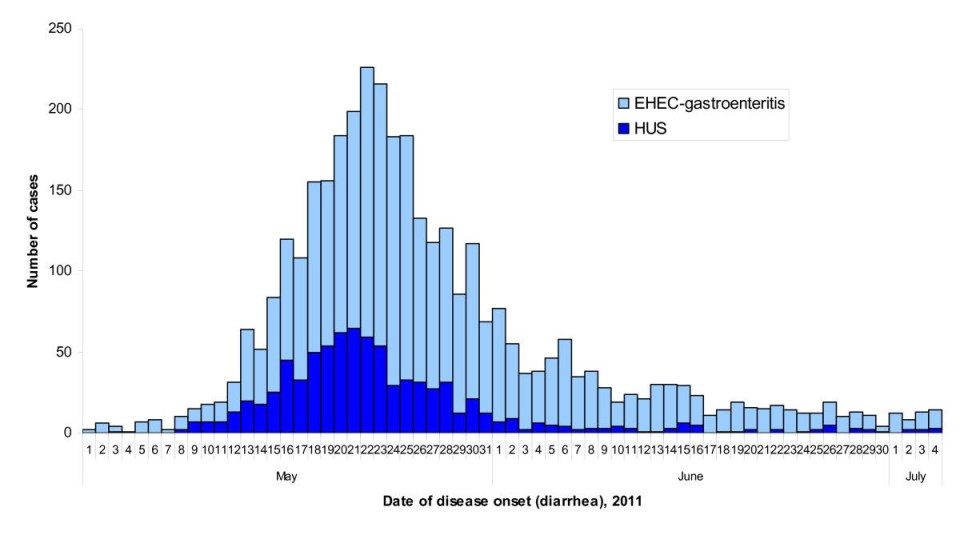 Figure 3: Epidemic curve of the 2011 outbreak of O104:H4 strain of Escherichia coli in Europe. Source: Werber D, Krause G, Frank C, Fruth A, Flieger A, Mielke M, Schaade L, Stark K. 2012. Outbreaks of virulent diarrheagenic Escherichia coli- are we in control? BMC Medicine. [accessed 2017 Nov 17]; https://bmcmedicine.biomedcentral.com/articles/10.1186/1741-7015-10-11. doi: 10.1186/1741-7015-10-11.
Figure 3: Epidemic curve of the 2011 outbreak of O104:H4 strain of Escherichia coli in Europe. Source: Werber D, Krause G, Frank C, Fruth A, Flieger A, Mielke M, Schaade L, Stark K. 2012. Outbreaks of virulent diarrheagenic Escherichia coli- are we in control? BMC Medicine. [accessed 2017 Nov 17]; https://bmcmedicine.biomedcentral.com/articles/10.1186/1741-7015-10-11. doi: 10.1186/1741-7015-10-11.
Aftermath
The outbreak had enormous economical consequences because the sale of salads and other vegetables decreased. Spanish vegetable producers were especially affected because Spanish cucumbers and tomatoes had been discussed as a potential source of pathogen. The European Union compensated farmers from several vegetable-exporting countries for a total of 220 million Euros.
Many criticized how the German government dealt with the outbreak. The notification time from the local health department to the regional and national agencies could have been shortened by a few days. Since then, Germany has put in place a centralized database shared by all health authorities to allow a faster parallel communication between all agencies.
References
Altmann M, Wadl M, Altmann D, Benzler J, Eckmanns T, Krause G, Spode A, An der Heiden M. 2011. Timeliness of Surveillance during Outbreak of Shiga Toxin–producing Escherichia coli Infection, Germany, 2011. Emerging Infectious Diseases. [accessed 2017 Nov 17]; 17(10): 1906-1909. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3310688/. doi : 10.3201/eid1710.111027.
Bielaszewska M, Mellmann A, Zhang W, Köck R, Fruth A, Bauwens A, Peters G, Karch H. 2011. Characterisation of the Escherichia coli strain associated with an outbreak of haemolytic uraemic syndrome in Germany, 2011: a microbiological study. The Lancet Infectious Diseases. [accessed 2017 Nov 17]; 11(9): 671-676. http://www.sciencedirect.com/science/article/pii/S1473309911701657. doi: 10.1016/S1473-3099(11)70165-7.
Burger R. 2012. EHEC O104:H4 in Germany 2011: Large Outbreak of BloodyDiarrhea and Haemolytic Uraemic Syndrome by Shiga Toxin-Producing E. Coli via Contaminated Food. In: Improving Food Safety Through a One Health Approach: Workshop Summary. Washington (DC): National Academies Press; [accessed 2017 Nov 17]. https://www.ncbi.nlm.nih.gov/books/NBK114499/.
Centers for Disease Control and Prevention (CDC). 2013. Outbreak of Escherichia coli O104:H4 Infections Associated with Sprout Consumption — Europe and North America, May–July 2011. Morbidity and Mortality Weekly Report (MMWR). [updated 20 December 2013; accessed 17 November 2017]; 62(50);1029-1031. https://www.cdc.gov/mmwr/preview/mmwrhtml/mm6250a3.htm.
Frank C, Werber D, Cramer JP, Askar M, Faber M, An der Heiden M, Bernard H, Fruth A, Prager R, Spode A, et al. 2011. Epidemic Profile of Shiga-Toxin–Producing Escherichia coli O104:H4 Outbreak in Germany. The New England Journal of Medicine. [accessed 2017 Nov 17]; 365: 1771-1780. http://www.nejm.org/doi/full/10.1056/NEJMoa1106483#t=article. doi : 10.1056/NEJMoa1106483.
Karch H, Denamur E, Dobrindt U, Finlay BB, Hengge R, Johannes L, Ron EZ, Tonjum T, Sansonetti J, Viencente M. 2012. The enemy within us: lessons from the 2011 European Escherichia coli O104:H4 outbreak. EMBO Molecular Medicine. [accessed 2017 Nov 17]; 4(9): 841-848. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3491817/. doi : 10.1002/emmm.201201662.
Youn Lim J, Yoon JW, Hovde CJ. 2010. A Brief Overview of Escherichia coli O157:H7 and Its Plasmid O157. Journal of Microbiology Biotechnology. [updated 20 January 2010; accessed 17 November 2017]; 20(1):5–14. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3645889/. doi: PubMed Central PMCID: PMC3645889.
